Dou et al., 2020 MIST: A multiloci identification system for Trichoderma AEM
Abstract
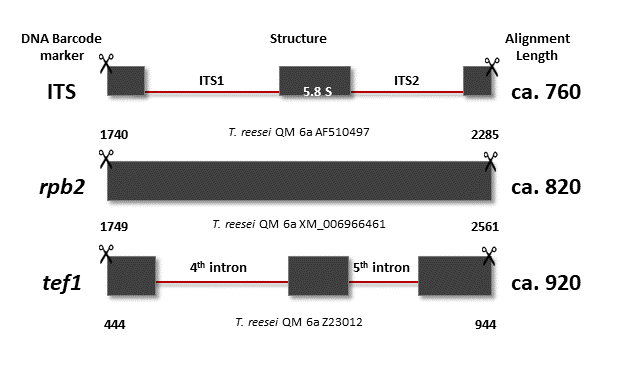
Due to the rapid expansion in microbial taxonomy, precise identification of common industrially and agriculturally relevant fungi such as Trichoderma species is challenging. In this study, we introduce the on-line multiloci identification system (MIST) for automated detection of three hundred and forty-nine Trichoderma species based on a set of three DNA barcodes. MIST is based on the reference databases of validated sequences of three commonly used phylogenetic markers collected from public databases. The databases consist of four hundred and sixteen complete sequences of the nuclear rRNA internal transcribed spacers 1 and 2 (ITS), five hundred and eighty-three sequence fragments of the gene encoding translation elongation factor 1-alpha (tef1) and five hundred and thirty-four sequence fragments of the gene encoding RNA polymerase subunit 2 (rpb2). Through MIST, information from different DNA barcodes can be combined and the identification of Trichoderma species can be achieved based on the integrated parametric sequence similarity search (blastn) performed in the manner of a decision tree classifier. In the verification process, MIST provided correct identification for forty-four Trichoderma spp. based on DNA barcodes consisting from tef1 and rpb2 markers. Thus, MIST can be used to obtain an automated species identification as well as to retrieve sequences required for the manual identification by means of phylogenetic analysis.
IMPORTANCE The genus Trichoderma is important to human kind, with a wide range of applications in industry, agriculture and bioremediation. Thus, quick and accurate identification of Trichoderma species is paramount since it is usually the first step in Trichoderma-based research. However, it frequently becomes a limitation, especially for researchers who lack taxonomic knowledge of fungi. Moreover, as the number of Trichoderma-based studies has increased, a growing number of unidentified sequences have been stored in public databases, which has made the species identification more ambiguous. In this study, we provide an easy to use tool for automated species identification (MIST), a list of Trichoderma species, and corresponding sequences of reference DNA barcodes. Therefore, this study will facilitate the research on the biodiversity and applications of the genus Trichoderma.



No comments! Be the first commenter?